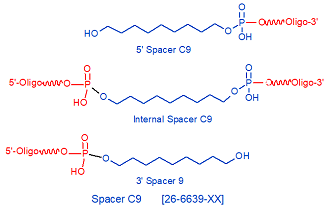
Modification : Spacer C9
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
| Catalog No | Scale | Price |
| 26-6639-05 | 50 nmol | $162.00 |
| 26-6639-02 | 200 nmol | $162.00 |
| 26-6639-01 | 1 umol | $216.00 |
| 26-6639-03 | 2 umol | $270.00 |
| 26-6639-06 | 5 umol | $972.00 |
| 26-6639-10 | 10 umol | $1,728.00 |
| 26-6639-15 | 15 umol | $2,160.00 |
| Discounts are available for Spacer C9! |
| Modification* Discount Price Structure |
|
1 site/order
|
List price
|
|
2 sites/order
|
10% discount
|
|
3 sites/order
|
20% discount
|
|
4 sites/order
|
30% discount
|
|
5-9 sites/order
|
50% discount
|
|
10+ sites/order
|
60% discount
|
|
*Exceptions apply
|
Spacer modifications C2, C3, C4, C6, C9, C12 and triethylene glycol Spacer 9 (PEG30 and 18 (PEG6)are used to insert a spacer arm in an oligonucleotide. These modifications can be added in multiple additions when a longer spacer is required. 3'-Spacer C3 CPG may also act as a blocker of exonuclease and polymerase activity at the 3'-terminus. dSpacer is used to introduce a stable abasic site within an oligonucleotide. See "Photo-Cleavable" modification category for photo-cleavable Spacer.
Spacer C9 is a 9-carbon spacer that is used to incorporate a long spacer arm into an oligonucleotide. Spacer C12 can be incorporated in consecutive additions if a longer spacer is required. Spacers are frequently used for solid-phase immobilization of DNA probes or aptamers for microarray applications (1,2), but can be used for any oligonucleotide-based application requiring a long spacer arm.
References
1. Reese, M.O., van Dam, R.M., Scherer, A., Quake, S.R. Microfabricated Fountain Pens for High-Density DNA Arrays.
Genome Res. (2003),
13: 2348-2352.
2. Lao, Y-H., Peck, K., Chen, L-C. Enhancement of Aptamer Microarray Sensitivity through Spacer Optimization and Avidity Effect.
Anal. Chem. (2006),
81: 1747-1754.
Oligonucleotide PEGylation : Spacers vs. PEGylation
Gene Link offers short PEG3 and PEG6 as direct coupling using automated chemistry. The PEG3 is termed as Spacer 9 and PEG6 as spacer 18. These are also used to introduce space between adjacent sequence and modifications. These can be inserted multiple times to increase the PEG units.
Larger 2, 5, 10 and 20 kDa PEGylation of oligonucleotides is inserted at any site of an oligonucleotide using a post synthesis amino group on the oligo with PEG-NHS.
PEGylation is the covalent attachment of polyethylene glycol (PEG) to oligonucleotides such as DNA, RNA, antisense, siRNA and aptamers. It improves pharmacokinetics, reduces renal clearance, increases nuclease stability, and decreases immunogenicity. (1) The way PEG shields its conjugated payload offers new challenges and opportunities for oligonucleotide PEGylation. Other than aptamers, the target of most oligonucleotides is a complementary sequence.
Messenger RNA (mRNA) delivery strategies are required to protect biologically fragile
mRNA from ribonuclease (RNase) attacks to achieve efficient therapeutic protein expression. To tackle this issue, most mRNA delivery systems have used cationic components.
A cation-free delivery strategy by hybridization of PEGylated RNA oligonucleotides with mRNA. The PEG strands on the mRNA sterically and electrostatically shields the mRNA, improving mRNA
nuclease stability 15-fold and the PEGylated mRNA induced nearly 20-fold higher efficiency of reporter protein expression than unhybridized mRNA in cultured cells (2).
PEGylation has been used to improve the biopharmaceutical properties of protein drugs since the 1990s, and over a dozen PEGylated pharmaceuticals are currently on the market (2). PEG creates a large hydration shell, which sterically blocks other biomacromolecules from penetrating through the polymer layer and binding with the interior substrate (3, 4). Binding requires displacing the PEG by the incoming molecule, generally making such binding less thermodynamically favorable. These properties usually result in weaker interactions between the receptor and the conjugated molecule, but increased drug solubility, prolonged blood circulation, and increased drug stability often offset by the reduced binding affinity. PEGylated oligonucleotides can be an exception to this generalization, with increased binding to a complementary sequence compared to unmodified ONs. The effect is attributed to macromolecular volume exclusion (6).
PEGylation References
1. Li WJ; Zhan P; De Clercq E; Lou HX; Liu XY Current drug research on PEGylation with small molecular agents. Prog. Polym. Sci 2013, 38, 421-444.
2. Yoshinaga, N; Naito, M; Tachihara, Y; Boonstra, E; Osada, K; Cabral, H and Uchida, S. PEGylation of mRNA by Hybridization of Complementary PEG-RNA Oligonucleotides Stabilizes mRNA without Using Cationic Materials. Pharmaceutics 2021, 13, 800.
3. Harris JM; Chess RB Effect of pegylation on pharmaceuticals. Nat. Rev. Drug Discov 2003, 2, 214-221. [PubMed: 12612647]
4. Harris JM; Martin NE; Modi M Pegylation. Clin. Pharmacokinet 2001, 40, 539-551. [PubMed: 11510630]
5. Plesner B; Fee CJ; Westh P; Nielsen AD Effects of PEG size on structure, function and stability of PEGylated BSA. Eur. J. Pharm. Biopharm 2011, 79, 399-405. [PubMed: 21620970]
6. Nakano S-I; Karimata H; Ohmichi T; Kawakami J; Sugimoto N The effect of molecular crowding with nucleotide length and cosolute structure on DNA duplex stability. J. Am. Chem. Soc 2004, 126, 14330-14331. [PubMed: 15521733]
- Spacer C9
