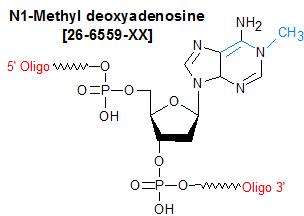
Modification : N1-Methyl dA (m1dA)
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
26-6559
Structural Studies
[m1dA]
Y
Y
Y
328.24
15.4
PS26-6559.pdf
-
-
-
| Catalog No | Scale | Price |
| 26-6559-05 | 50 nmol | $324.00 |
| 26-6559-02 | 200 nmol | $324.00 |
| 26-6559-01 | 1 umol | $421.00 |
| 26-6559-03 | 2 umol | $632.00 |
| 26-6559-06 | 5 umol | $1,894.50 |
| 26-6559-10 | 10 umol | $3,370.00 |
| 26-6559-15 | 15 umol | $4,212.00 |
| Discounts are available for N1-Methyl dA (m1dA)! |
| Modification* Discount Price Structure |
|
1 site/order
|
List price
|
|
2 sites/order
|
10% discount
|
|
3 sites/order
|
20% discount
|
|
4 sites/order
|
30% discount
|
|
5-9 sites/order
|
50% discount
|
|
10+ sites/order
|
60% discount
|
|
*Exceptions apply
|
N1-Methyl-deoxyadenosine (N1-Me-dA, m1 dA) is a methylated nucleoside base, and is primarily used in the study of DNA damage and repair mechanisms related to alkylation damage. The N1-Me-dA lesion is primarily generated by SN2 alkylating reagents such as methyl methanesulfonate and dimethylsulfate, which react with the N1 position of adenine (1). In cells, N1-methyl-dA acts as a lethal DNA replication block, but is not very mutagenic (1% A to T transversion in E. coli), and is repaired by the enzyme AlkB by direct reversal (2,3). Because the N1 position of adenine is involved in hydrogen bonding of A : T Watson-Crick base pairing, methylation of this site was expected to disrupt hydrogen bonding. However, NMR analysis revealed that N1-methylation actually alters the A:T base-pairing interactions from Watson-Crick to (syn)N1-methyl-A : (anti)T Hoogsteen, thus providing insight into why AlkB repair of N1-Methyl-dA lesions is 10X more efficient on ssDNA over dsDNA (4).
References
(1) Sedgwick, B., Lindahl, T. Recent progress on the Ada response for inducible repair of DNA alkylation damage.
Oncogene (2002),
21: 8886-8894.
(2) Chen, B.J., Carroll, P., Samson, I. The Eschericia coli alkB protein protects human cells against alkylation-induced toxicity.
J. Bacteriol. (1994),
176: 6255-6261.
(3) Delaney, J.C., Essigman, J.M. Mutagenesis, genotoxicity and repair of 1-methyladenine, 3-alkylcytosines, 1-methylguanine, and 3-methylthymine in alkB Escherichia coli.
Proc. Natl. Acad. Sci. (USA) (2004),
101: 14051-14056.
(4) Yang, H., Zhan, Y., Fenn, D., Chi, L.M., Lam, S.K. Effect of 1-methyladenine on double-helical DNA structures.
FEBS Letters (2008),
582: 1629-1633.
- N1-Methyl dA (m1dA)
