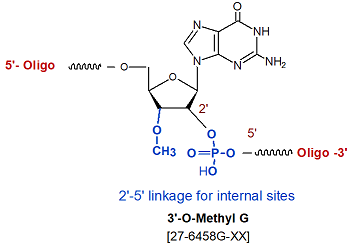
Modification : 3'-O methyl rG
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
| Catalog No | Scale | Price |
| 27-6458G-05 | 50 nmol | $227.00 |
| 27-6458G-02 | 200 nmol | $227.00 |
| 27-6458G-01 | 1 umol | $297.00 |
| 27-6458G-03 | 2 umol | $335.00 |
| 27-6458G-06 | 5 umol | $1,336.50 |
| 27-6458G-10 | 10 umol | $2,376.00 |
| 27-6458G-15 | 15 umol | $2,970.00 |
| Discounts are available for 3'-O methyl rG! |
| Modification* Discount Price Structure |
|
1 site/order
|
List price
|
|
2 sites/order
|
10% discount
|
|
3 sites/order
|
20% discount
|
|
4 sites/order
|
30% discount
|
|
5-9 sites/order
|
50% discount
|
|
10+ sites/order
|
60% discount
|
|
*Exceptions apply
|
Antisense Oligos (ODN) & siRNA Oligo Modifications
Click here for more information on antisense modifications, design & applications.
3'-O methyl bases (3'-Ome) form a 2'-5' phosphodiester linkage when placed internally. A single 3'-O methyl base modification at the 3' end will have a hydroxyl group at the 2' end and 3'-O methyl at the 3' end, this will prevent the oligo from extension by polymerases.
3'-O methyl bases (3'-Ome)-(2'-5' linked), are used to substitute 2'-5' phosphodiester linkages for the usual 3'-5' phosphodiester linkages at some or all positions of an oligonucleotide. Oligonucleotides containing all, or primarily, 2', 5'- phosphodiester linkages selectively bind to complementary single-stranded 3', 5'- RNA over comparable 3', 5'- DNA. Presumably this selectively is a consequence of the 2', 5'- linkages destabilizing duplexes formed with 3', 5'- DNA more than those formed with 3', 5'-RNA, leading to 2'-5'- RNA:3', 5'- DNA duplexes having much lower Tm than the corresponding 2'-5'- RNA:3'- 5'-RNA duplexes. This property means that RNA oligos containing such linkages could be useful in anti-sense applications, as ssRNA-specific probes, or as ligands for affinity purification of cellular RNA.
Antisense oligonucleotides (ASOs) and small interfering RNA (siRNA) are both recognized therapeutic agents for the silencing of specific genes at the posttranscriptional level. Chemical modifications, particularly 2'-O-(2-Methoxyethyl)- oligoribonucleotides (2'-O-MOE bases) and 2'-OMethyl bases are commonly used to confer nuclease resistance to an oligo designed for anti-sense, siRNA or aptamer-based research, diagnostic or therapeutic purposes, when specific 2'-OH is not required.
Nuclease resistance can be further enhanced by phosphorothiolation of appropriate phosphodiester linkages within the oligo. These modifications confers nuclease resistance, high binding affinity towards complementary RNA, reduced unspecific protein binding and extended half-life in tissues.
Gapmers. Currently, the mainstream of the ASO is gapmer design ASOs. Gapmer design oligonucleotides, contain two to
five chemically modified nucleotides (LNA, 2'-O methyl or 2'-O-MOE RNA) as 'wings' at each terminus flanking a central 5- to 10-base 'gap' of DNA, enable cleavage of the target mRNA by RNase H, which recognizes DNA/RNA heteroduplexes. Usually all the phosphodiester linkages are converted to phosphorothioate.
Delivery. The development of effective delivery systems for antisense oligonucleotides is essential for their clinical therapeutic application. The most common delivery system involves a relatively hydrophobic molecule that can cross the lipid membrane. The following list of modifications are suitable for delivery system in addition to cell penetrating peptides.
Cholesterol
Tocopherol (alpha-tocopherol, a natural isomer of vitamin E)
PEG
- 3'-O methyl rG (2'-5' linked)
