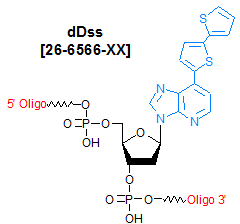
Modification : dDss
Catalog Reference Number
Category
Modification Code
5 Prime
3 Prime
Internal
Molecular Weight (mw)
Extinction Coeficient (ec)
Technical Info (pdf)
Absorbance MAX
Emission MAX
Absorbance EC
26-6566
Duplex Stability
[dDss]
N
N
N
461.45
8.4
PS26-6566.pdf
-
-
-
| Catalog No | Scale | Price |
| 26-6566 | 50 nmol | $0.00 |
| 26-6566 | 200 nmol | $0.00 |
| 26-6566 | 1 umol | $0.00 |
| 26-6566 | 2 umol | $0.00 |
| 26-6566-06 | 5 umol | $1,894.50 |
| 26-6566 | 10 umol | $0.00 |
| 26-6566 | 15 umol | $0.00 |
This modification is discontinued. Consider dNaM [26-6561] and d5SICS [26-6562] as substitutes. See related products.
Base Pair Recognition Through Hydrophobic Interactions
The unnatural base pair between 7-(2-thienyl)-imidazo[4,5-b]pyridine (Ds) and pyrrole-2-carbaldehyde (Pa) is formed by specific hydrophobic shape complementation. The shape of the Ds-Pa pair is different from those of the natural A-T and G-C pairs, but the Ds-Pa pair works together with the natural pairs in in vitro replication and transcription. Pa also functions as a template base for incorporating another unnatural base, 2-amino-6-(2-thienyl)purine (s), into RNA. The s base also acts as a unique fluorescent base analog in DNA and RNA fragments. dDss is strongly fluorescent and is useful as a fluorescent tag for DNA detection. dDss also forms a base pair with dPa. Biotin PaTP can be site-specifically incorporated into RNA, opposite dDs at a desired position in DNA templates, by T7 transcription. Similarly, the fluorescent s base can be site-specifically incorporated into RNA opposite dPa in DNA templates.
The dNaM and d5SICS matched pair appears to be novel base pair. These unnatural C-nucleosides have pair recognition that rivals the A-T and G-C pairing in the natural genetic alphabet. In addition,
they have been shown to be well-replicated by DNA polymerases under steady-state conditions,
regardless of sequence. The fidelity and efficiency of dNaM and d5SICS replication approach
those of natural synthesis. Both dNaM and d5SICS are also efficiently transcribed by T7 RNA
polymerase in either direction.
See Glen Report for details:
Unnatural Bases
References
1. Seo, Y.J.; Matsuda, S.; Romesberg, F.E. J. Am. Chem. Soc., 2009, 131(14), 5046-7.
2. Seo, Y.J.; Hwang, G.T.; Ordoukhanian, P.; Romesberg, F.E. J. Am. Chem. Soc., 2009,
131(9), 3246-52.
3. Seo, Y.J.; Romesberg, F.E. ChemBioChem, 2009, 10(14), 2394-2400.
4. Hwang, G.T.; Hari, Y.; Romesberg, F.E. Nucleic Acids Res., 2009, 37(14), 4757-63.
5. Hari, Y.; Hwang, G.T.; Leconte, A.M.; Joubert, N.; Hocek, M.; Romesberg, F.E.
ChemBioChem, 2008, 9(17), 2796-9.
6. Hwang, G.T.; Romesberg, F.E. J. Am. Chem. Soc., 2008, 130(44), 14872-82.
7. Leconte, A.M.; Hwang, G.T.; Matsuda, S.; Capek, P.; Hari, Y.; Romesberg, F.E. J. Am.
Chem. Soc., 2008, 130 (7), 2336-43.
- dDss
